The new age of brain maps
The laboratory of Prof George Paxinos is using technological advances in brain imaging to create new atlases of the anatomy of the nervous system.
Brain researchers, no less than geographers, need maps and coordinate systems to navigate the brain and communicate their observations to each other. Think of a Google map depicting the shape of an island or the streets in a city. Just as this map can be overlaid with bicycle routes, train stations, or elevation data, a simple map of the brain can be superimposed by types of neurons, neurotransmitters, enzymes, connectivity, and functional data.
In brain imaging research, an atlas, or collection of brain maps, is typically understood as a reference frame or coordinate system into which a newly acquired dataset is transformed. This provides a numerical reference frame to share findings, but also to infer what anatomical detail or structure is expected at a given coordinate.
The Paxinos lab at NeuRA is now constructing the new generation of brain and spinal cord maps to assist in studying disease, interpreting research imaging data, and improving the accuracy of therapeutic interventions on the human brain (e.g., surgery).
A high resolution MRI atlas (a collaboration between George Paxinos and Charles Watson)
By far the most widely used animal brain atlas for studying animal models of human brain pathology is the histological atlas of the rat brain of Paxinos and Watson. This atlas identifies over 650 nuclei, or brain structures, and 140 pathways and has been crucial to the success of many thousands of studies. It is the most widely cited publication in all of neuroscience, with 61,376 citations.
Paxinos and Watson are now producing the first of a new generation of atlases exploiting the combined power of three neuroscientific and computing techniques: magnetic resonance histology, diffusion tensor imaging, and digital atlasing. This project is in collaboration with Al Johnson at Duke University and supported by the NHMRC and the ARC Centre of Excellence for Integrative Brain Function.
Accurate magnetic resonance (MR) atlases will have many practical and technical advantages over a histological atlas. They can generate 3D images and can be used to guide studies of living animals that can be scanned repeatedly without injury. Conventional histological procedures will continue to provide the highest resolution atlases for two-dimensional planar images. However, these atlases suffer significant distortion arising from tissue processing. The resolution of the MR data in the images acquired for this project is sufficiently high that traditional two-dimensional histology sections can be registered, or matched, to it. This MR atlas is expected to have an impact on studies of rodent models of human brain disease.
Making technological strides
Earlier attempts to produce an MR atlas of the rat suffered from various technical limitations. Super high field (7 Tesla (T)) MRI systems (human MRI machines are usually 1.5 to 3T) now allow resolutions that were unthinkable until just recently. The large arrays used in the present project (800 x 800 x 1600 Angstroms (Å)) represent an increase in data volume of more than 30 times that of older rat atlases.
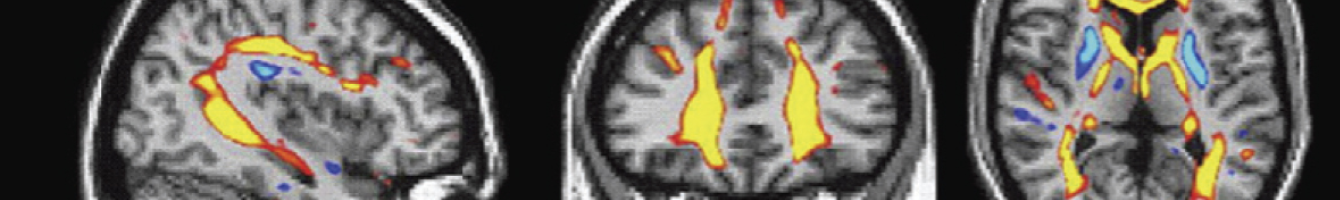
Using a high field MRI system a number of data extraction sequences have been developed, which emphasize different contrasts in the tissue. These contrasts show different aspects of the anatomy of the brain, in the same way as conventional histology atlases have made use of a range of histochemical marker stains. The most important of these extraction sequences have been gradient recalled echo (GRE), diffusion weighted image (DWI), apparent diffusion coefficient (ADC), radial diffusivity (RD), and color-encoded fractional anisotropy (FAC).

Brain sections with various contrasts applied that each highlight anatomical characteristics in a different way. L-R: cresyl violet stain, acetylcholinesterase stain, final published diagram, pencil delineations on a gradient-recalled echo (GRE) scan, diffusion weighted image (DWI) scan, and a colour-encoded fractional anisotropy (FAC) scan
Analog methods play a role too
The first step in adding the detail to each brain image is the preparation of a master set on which anatomical delineations of nuclei and pathways can be made. In these image sets the major outlines of the parts of brain are clear and easy to trace. A2-size photographs of all GRE images (approximately 100) are printed and covered with thin mylar transparency paper. All annotations are made on these transparency covers. It may seem odd to choose to work in this way, when the images are digital and can even be displayed simultaneously in the three cardinal planes (coronal, sagittal and horizontals). However, the experience of Paxinos and Watson in making many brain atlases is that it is far more efficient to work with paper overlay than with on-screen images.
They have looked hard at on-screen alternatives, but find them clumsy and slow for the preparation of the initial draft outlines and labels. Of course, as soon as the draft and labelled outlines are complete, they convert them to Adobe Illustrator files, and all subsequent corrections are done digitally. The team will attempt to identify all the major nuclei and tracts on the basis of data from all five contrasts.
The human brainstem in 3D (a collaboration between George Paxinos and Mark Schira)
Another new brain mapping development is the creation of an atlas focusing on the human brainstem. The brainstem is critical for vital functions such as breathing, movement and arousal, and cognition and emotion. Different parts of the brainstem have been reported to be involved in a range of human pathologies, including progressive supranuclear palsy, multiple system atrophy, autism spectrum disorders, and Parkinson’s disease. Until recently, the human brainstem was largely inaccessible to in vivo MR imaging because many of the structures were too small and the MRI contrast too low. Technical advances have now dramatically improved in vivo imaging of the brainstem. An electronic 3D atlas of the human brainstem being constructed in the Paxinos laboratory will combine three data sources – in vivo MRI data, MR microscopy data, and finally, histology data – into a common 3D space that allows switching between the various data types. This work is supported by a project grant from the NHMRC.
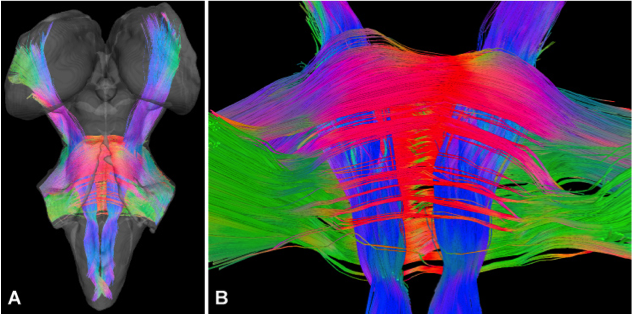
Brainstem tractography reconstruction based on human post-mortem DTI images. These images were created by Evan Calabrese and Al Johnson of Duke University.